Prognostic and predictive biomarkers in MPM: a new era?
Introduction
Malignant pleural mesothelioma (MPM) behaves in an aggressive manner and the current therapeutic approaches are generally unable to change the bleak prognosis that characterizes cancer patients with this disease. Moreover, it is important to keep in mind that the life expectancy of MPM is defined by known prognostic factors such as stage and histologic classification. Nevertheless, there are no validated predictors of the treatment response or of long-term outcomes, such as overall survival or survival free of disease progression. Many studies have examined ideal biomarkers as a predictive or prognostic approach to MPM. These include substances present in both the pleural fluid and in the blood to validate whether a minimally invasive analysis of these elements could provide early information about the disease.
It is well known that one of the most studied markers in pleural effusion is mesothelin, which shows good specificity but low sensitivity, especially for non-epithelial MPM. Other biomarkers found in pleural fluid include fibulin-3, hyaluronan, microRNAs, and CYFRA-21.1, which have lower diagnostic capabilities compared with mesothelin but provide prognostic information and play a potential role as therapeutic targets.
Serum is the most widely studied matrix for MPM biomarkers.
To this regard, several serum biomarkers of PM have been studied, of which mesothelin, osteopontin, and fibulin-3 are the ones most frequently tested. Soluble mesothelin-related peptide (SMRP) is the only FDA-approved biomarker in patients with suspected mesothelioma to provide useful information regarding diagnosis, prognosis, follow-up, and response to therapy in epithelial MPM. As far as future prospects are concerned, panels combining different markers and proteomic technologies look promising in terms of improving clinical performance in the diagnosis and monitoring of the disease under specific treatment. In this scenario, one might glimpse the dawn of a new era: obtaining important information about the disease and the patient with MPM through inexpensive, repeatable, and minimally invasive analyses: the discovery of ideal prognostic and predictive biomarkers.
Which biomarkers are available today?
The search for biomarkers useful for MPM has been of great interest for some time now, with three main potential purposes:
1) Screening in people at risk (both those exposed to asbestos and family members in cases of "genetically related" MPM);
2) Improvement of the diagnostic process in patients with pleural effusion or other abnormalities, such as nonspecific pleural thickening;
3) Evaluation of the treatment response and prognostic evaluation.
In this context, the detection of soluble or pleural fluid biomarkers of MPM could be useful to cut down on the need for invasive procedures in patients with poor performance status.
The 2018 British Thoracic Society's guidelines for investigating and managing MPM recommend that biomarker analysis be taken into consideration only in patients with suspicious cytology who are not sufficiently suitable for more invasive diagnostic tests, whereas these biological predictors are not suggested for screening or diagnosis of MPM, nor for predicting treatment response or survival [16].
For the management of MPM, the European Respiratory Society (ERS/European Society of Thoracic Surgery (ESTS)/European Association of Cardio-Thoracic Surgery (EACTS)/European Society of Radiotherapy and Oncology (ESTRO) 2020) does not suggest routine determination of mesothelin or other biomarkers for diagnosis, screening, or prognostic assessment, in the absence of clear scientific evidence as to their usefulness [17].
Screening methods should be minimally invasive and cost-effective tests capable of identifying MPM in the population of asbestos-exposed individuals so as to potentially treat the disease at an early stage.
Although it would certainly be desirable to have very sensitive and specific tests, high specificity is strongly recommended for screening asymptomatic subjects to reduce the number of false positives as well as the resulting unnecessary procedures and psychological stress for the subjects involved [18]. High specificity is also necessary for tests that concern the differential diagnosis of pleural lesions, most of which are obtained by pleural biopsies. The complication rate, morbidity, and costs associated with pleural biopsies depend on the procedure used to obtain the specimen, such as medical thoracoscopy, thoracic surgery, and transthoracic image-guided biopsy [19].
The diagnostic definition of MPM and its distinction from nonmalignant lesions are crucial when making management decisions involving the patient and family members.
Though relatively invasive, medical or surgical thoracoscopy can allow diagnosis and staging and, at the same time, offer permanent treatments to counteract the onset of malignant pleural effusion (e.g., indwelling placement of a pleural catheter and/or palliative pleurodesis) [20].
In the past two decades, several studies have investigated the diagnostic and prognostic markers of MPM, but as of today, the most reliable predictors of disease outcome are still clinical and pathological parameters. In particular, non-epithelial histology, poor performance status, male sex, anemia, thrombocytosis, leukocytosis, high LDH, advanced age, and advanced disease are unfavorable prognostic indicators in patients with MPM. Many of these parameters are included in the most widely used scoring systems for MPM, namely that of the European Organization for Research and Treatment of Cancer (EORTC) and Cancer and Leukemia Group B (CALGB). They were introduced more than 20 years ago and are still valid, although many studies have attempted to update them, identify additional prognostic biomarkers, and develop models that combine clinical and molecular characteristics [21, 22].
Candidate biomarkers can be molecules with different traits, such as proteins or their fragments, nucleic acids, lipids and metabolites. As a result, methods for their identification vary considerably.
Proteomics-based approaches, such as those based on mass spectrometry, represent promising tools and have been increasingly implemented to identify and quantify biomolecules in a variety of biological samples.
Nucleic acid evaluation is more complex, as it requires RNA extraction, quantification and purification, with samples stored at very low temperatures (-80 °C). The purified RNA is then transcribed in reverse into cDNA, which, in turn, is amplified through polymerase chain reaction (PCR).
Nucleic acid evaluation and, even more so, proteomics are expensive and present problems in terms of accessibility in low-income countries.
Pleural biomarkers
Many molecules present in pleural fluid have been studied as potential biomarkers of malignancy.
In particular, pleural fluid biomarkers are potentially promising diagnostic tools for MPM, which in about half of cases manifests itself as pleural effusion. However, the diagnostic value of the most studied biomarkers for MPM in pleural effusion is still largely to be defined [17].
In this regard, the greatest attention of the scientific community has been focused on mesothelin and mesothelin-related soluble peptides (SMRPs), fibulin-3, osteopontin, and cell-free microRNAs.
In a recent meta-analysis on PM markers, 36% of the included studies searched for them in pleural effusions. Proteins were the most studied biomarkers (89%), while DNA and miRNAs were studied in only 5.5% of the studies [25].
MESOTHELIN
Mesothelin is a protein usually found in the mesothelial cells of the pleura, peritoneum, and pericardium. It appears to play a role in cell adhesion but is probably a nonessential component in normal cells.
Mesothelin is overexpressed in some malignancies, such as MPM, but also in adenocarcinomas of the pancreas, ovary, and lung [47]. Therefore, this protein may be considered an interesting biomarker for the diagnosis of MPM, for screening people exposed to asbestos, for monitoring disease progression, and as a potential target for cancer therapy [48].
Several pioneering studies on mesothelin found that individuals with the epithelial subtype had mesothelin expression, whereas those with the sarcomatous subtype were all negative for this biomarker; moreover, in patients with biphasic MPM, only the epithelial component showed positivity for mesothelin [50] [51] [52]. Consequently, positive immunostaining for mesothelin suggests epithelial MPM, although it is not absolutely specific.
The meta-analyses by Gao et al. [26] and Cui et al. [27] evaluated the diagnostic accuracy of SRMPs on pleural fluid for the diagnosis of MPM. They analyzed SMRP concentrations in 13 and 11 studies, respectively, and all reported higher levels of mesothelin in MPM patients compared with controls. Overall, the authors concluded that MPM can be suspected when SMRPs in pleural effusion are above 8 nmol/L, while values above 20 nmol/L are strongly suggestive.
A more recent meta-analysis by Schillebeeckx et al. [25] included 19 studies that evaluated the diagnostic efficacy of mesothelin in pleural effusion. Cut-off values ranged from 6 nM (where sensitivity and specificity were 72% and 46%, respectively) to 6 nM.
FIBULIN-3
Fibulin-3 is a glycoprotein encoded by the fibulin-like extracellular matrix protein 1 gene containing the epidermal growth factor. It plays a role in cell proliferation and migration [53, 54].
Fibulin-3 has low expression in normal tissues but is overexpressed in several tumors, including MPM, and is also secreted into body fluids. It accumulates in the pleural effusions of patients with MPM and has been suggested to distinguish these patients from individuals with nonmalignant pleural inflammation [28]. These findings were not confirmed by other studies that found similar fibulin values in effusions from mesothelioma and other diseases [55].
A comparative analysis suggested that fibulin-3 correlates less precisely than mesothelin with the diagnosis of PM, whether measured in plasma or pleural effusion; consequently, mesothelin has been recognized as the best pleural marker that can be used for routine diagnostic purposes [56, 57]. Conversely, fibulin-3 has been put forward as a better prognostic factor than PM; genuine evidence suggests that fibulin-3 promotes malignant behavior of mesothelial cells, while the knockdown of fibulin-3 decreases the viability, clonogenic capacity and invasion, as well as the chemoresistance, in PM cells.
HYALURONIC ACID
Hyaluronan, or hyaluronic acid (HA), is a polysaccharide that seems to be involved in the progression of several types of neoplasms [58]. Several scientific studies have demonstrated high levels of HA in pleural effusions associated with mesothelioma [29], due to the release of growth factors from tumor cells that can stimulate other cells to produce HA [59]. HA is rapidly removed from the circulation by the stabilin-2 clearance receptor and has a plasma half-life of 2.5-5 minutes [29]. In the past, measuring hyaluronic acid by high-performance liquid chromatography required high technical expertise, which has limited the number of studies on this biomarker.
More recent research, using faster analytical systems within the reach of any scientific laboratory, has shown that mesothelin and hyaluronic acid levels in pleural effusion have similar diagnostic accuracy and that combining the two markers could improve diagnostic accuracy [60].
MicroRNAs
MicroRNAs are short single-stranded noncoding RNA molecules that regulate gene expression at the post-transcriptional level. These molecules affect the course of many important processes in the human organism, including cell division, proliferation, differentiation, apoptosis, and blood vessel formation.
Some studies have evaluated serum levels of different microRNAs as markers of malignant mesothelioma, while searches for this biomarker in pleural fluid are rare [30]. The authors analyzed microRNAs in cells and supernatants of pleural effusion from 26 patients with MPM and 21 with pleural effusion caused to conditions other than MPM. The researchers found that four microRNAs (miR-944, miR-139-5p, miR-210 and miR-320) found in the pleural effusion were upregulated and seven (miR-200b, miR-200c, miR-143, miR-200a, miR-203, miR-31 and miR-874) were downregulated. A combination of miR-143, miR-210 and miR-200c was able to differentiate MPM from pleural effusions caused by other diseases.
CYFRA-21-1 and CEA
CYFRA-21-1 is the soluble fragment of cytokeratin 19. It can be released into the circulation after cell death, thus showing a close relationship with necrosis and apoptosis of cancer cells.
CYFRA-21-1 is found in the blood of patients with various epithelial malignancies, including non-small cell lung cancer (NSCLC), and has been used as a useful marker to predict diagnosis and prognosis [62].
Even though CYFRA-21-1 has not been extensively studied in MPM, all studies that have measured it in pleural effusion have found higher levels in patients with this neoplastic disease than in controls [31,63,64]. However, diagnostic accuracy was found to be modest.
CEA is a glycoprotein involved in cell adhesion.
In healthy individuals, very low levels of CEA are detectable in the bloodstream and body fluids, while its increase has been reported in several diseases including neoplastic conditions.
Two studies that measured CEA in pleural fluid reported its increase in MPM [31,65], while another research showed that pleural CEA in MPM was lower than in other types of cancer, suggesting that CEA levels above 3 ng/mL in pleural fluid may exclude the diagnosis of such pleural disease [64].
However, CEA currently shows poor diagnostic accuracy and is not recommended as a differential diagnostic biomarker for MPM [25].
COMBINING MARKER PANELS
In a study aimed at establishing a predictive model using biomarkers taken from pleural effusions, samples were collected from 190 consecutive patients [66].
The biomarkers significantly associated with MPM were hyaluronan, N-ERC/mesothelin, C-ERC/mesothelin, and syndecan-1.
Recently, a new proteomic method based on affinity-enrichment mass spectrometry was applied for the exploratory analysis of pleural effusions from a prospective cohort of 84 individuals undergoing thoracoscopy for suspected MPM [67]. The immunohistology of pleural biopsies confirmed MPM in 40 patients and ruled it out in 44 patients. The authors identified protein biomarkers with a marked ability to discriminate patients with MPM from those without MPM and applied a random forest algorithm in order to construct classification models.
Depending on the specific combination of proteins, the proteomic analysis of pleural effusions identified panels of proteins with excellent diagnostic properties (sensitivity of 90-100%, specificity of 89-98%, and AUCROC 0.97-0.99).
The proteins associated with cancer diagnosis include the galactin-3 binding protein, testican-2, haptoglobin, Beta ig-h3 and AMBP protein. Furthermore, the study confirmed the previously reported diagnostic accuracy of the two MPM markers fibulin-3 and mesothelin.
CYTOLOGY
The detection of neoplastic invasion has always been a key element in diagnosing MPM with certainty, but diagnosis based exclusively on the cytology of pleural effusion is controversial, mainly due to poor sensitivity [68]. The best interpretive output comes from correlating cytologic findings with imaging, which can provide information on the anatomic distribution of the lesion, evidence of nodularity of pleural disease, and sometimes tissue invasion.
Although "positive" and "negative" immunohistochemical markers have been shown to be remarkably effective in distinguishing epithelial mesothelioma from other secondary malignancies, no biomarker has 100% sensitivity or specificity for the diagnosis of mesothelioma. With regard to histology, MPM can be determined directly from morphology through hematoxylin-eosin staining. However, pathologists usually recommend confirmation via immunohistochemistry.
Calretinin, Wilms' tumor 1 (WT-1), cytokeratin 5 (CK5), podoplanin, mesothelin, and cardiac development protein with EGF-like 1 (HEG1) domains are immunohistochemical biomarkers of mesothelial differentiation, whereas Carcinoembryonic Antigen (CEA), B72. 3, Ber-EP4, Lewisy blood group (BG8), MOC-31, CD15, mucin-4 (MUC4) and claudin-4 are markers indicative of epithelial metastasis [2]. The loss of expression of the BAP1 protein by immunohistochemistry has recently been suggested as a potential marker for the identification of MPM since it has been observed in more than half of MPMs, both epithelial and biphasic or sarcomatoid [69].
Recent breakthroughs in cytologic analysis promise diagnostic advances for PMs. Recent studies suggest that cytologic evaluation of pleural effusions, assisted by HA evaluation, has a diagnostic accuracy for MPM that is not inferior to that of standard histologic evaluation and can be taken into consideration in difficult or unclear diagnostic cases.
Likewise, a large single-center database was retrospectively explored to shed light on the value of cytology in distinguishing malignant mesothelioma according to the International System for Reporting Serous Fluid Cytopathology (ISRSFC) [71]. Cytology samples from 210 patients with malignant mesothelioma (164 pleural effusions and 46 peritoneal effusions) were available for analysis. All cases were examined and reclassified according to the scheme proposed by the ISRSFC. The final histological diagnosis was epithelial mesothelioma in 192 patients (91.4%) and sarcomatoid-type mesothelioma in the remaining 18 patients (8.6%). The cytologic cases were reclassified as follows: 2 (0.9%) as nondiagnostic, 81 (38.6%) as formally negative, 4 (1.9%) as atypical of undetermined significance, 11 (5.2%) as formally suspicious, and 112 (53.4%) as malignant. Sarcomatoid cells in the malignant category appeared solitary, with moderate or marked nuclear pleomorphisms and irregular chromatin compared with the epithelial subtype. The authors concluded that morphologic features, together with clinical-radiologic data, can assist physicians in the proper management of patients.
Blood biomarkers
Serum is the most extensively studied matrix for MPM biomarkers; the most commonly studied biomarkers in recent research have been: mesothelin, osteopontin, and fibulin-3.
MESOTHELIN AND SMRP
Mesothelin has been the most widely studied serum biomarker in MPM [73]. A meta-analysis regarding the diagnostic value of soluble mesothelin in 4491 patients estimated the sensitivity and specificity of this marker to be 47% and 95%, respectively [74]. A more recent systematic review and meta-analysis, which included 27 studies, showed overexpression of the protein in patients with malignant mesothelioma.
Higher concentrations of this protein were detected in the epithelial type compared with other MPM subtypes [17, 75, 77-79].
Increased SMRP levels correlate with large tumor volume [80, 81].
Several studies have shown that longitudinal measurement can be used to assess tumor response and progression and can be associated with radiological findings [82-86]. However, some researchers have recently confirmed the correlation of SMRP and fibulin-3 with initial tumor volume, although more recent studies have not confirmed the clinical usefulness of biomarkers in terms of assessing tumor response in patients undergoing immunotherapy [81].
SMRP levels decrease after surgery and are a promising serum biomarker for detecting recurrence after the resection of epithelial MPM [73, 87].
Mesothelin expression represents a key criterion for selecting patients to undergo targeted treatments with mesothelin.
It has also been studied as a potential therapeutic target in patients with MPM.
OSTEOPONTIN
Serum osteopontin (OPN) shows good accuracy as regards the diagnosis of MPM when compared with healthy controls with or without asbestos exposure, but it is not useful when compared with benign pleural effusions [25,36] and other pathologies [37], thus suggesting the low specificity of this marker.
Plasma OPN shows higher accuracy compared with serum markers, as suggested by meta-analyses and head-to-head studies [25, 94-96]. This could be due to the easy degradation by thrombin in peripheral blood [97].
OPN may play a role in terms of prognosis. Several studies have shown that high OPN levels correlate with poor prognosis in patients with MPM [38-40, 95].
FIBULIN-3
The properties of plasma fibulin-3 as a marker were first reported by Pass et al. in 2012 [28]. In 507 patients from 3 cohorts, the authors found the biomarker to have a sensitivity and specificity of 95%, meaning that it might be useful in distinguishing MPM from asbestos-exposed healthy controls and patients with other malignancies.
These results were not confirmed by subsequent studies [57, 98-100], which showed lower accuracy of the biomarker.
However, a recent meta-analysis that included studies of plasma fibulin-3 showed the possible efficacy of plasma fibulin as a useful marker [25].
Head-to-head studies comparing the diagnostic performance of fibulin-3 and mesothelin/SMRP have shown conflicting and inconclusive results [36,57].
Several researches have demonstrated that serum/plasma fibulin-3 is not a reliable marker for either prognosis [38] or for assessing response to immunotherapy in MPM [81].
Fibulin-3 has also been suggested as a relevant molecular target to reduce the progression of MPM, and anti-fibulin-3 approaches are currently being investigated [88].
CALRETININ
Calretinin is a calcium-binding protein originally found in neurons but also expressed on the surface of mesothelial cells.
Calretinin is widely used in immunohistochemical evaluations of cytohistological specimens of suspected MPM, both epithelioid and sarcomatoid [32]. Its detection in plasma and serum does not differ significantly. Higher values of circulating calretinin have been detected in subjects with MPM compared with healthy controls exposed to asbestos [101].
Studies on mice primary mesothelial cells have suggested that overexpression of calretinin would promote the proliferation and migration of mesothelial cells [102]. Consequently, researchers have begun to speculate that calretinin may be a possible blood biomarker for screening as well as a new potential therapeutic target of MPM.
Studies have shown promising results for this marker in the early diagnosis of MPM and in differentiating MPM patients from asbestos-exposed and healthy patients [33, 103].
In addition, calretinin, both alone and in combination with mesothelin, was also evaluated in a large prospective cohort study of subjects with benign asbestos-related diseases who took part in annual screenings. The combination of the two markers obtained a sensitivity and specificity of 46% and 98%, respectively, in detecting mesothelioma up to about one year before the clinical diagnosis [104]
MicroRNAs (miRNAs) AND LONG NONCODING RNAs (lncRNAs)
The expression characteristics of miRNAs are associated with tumor type and clinical outcome, as demonstrated by genomic profiling. Therefore, miRNAs have a potential role as candidates for diagnostic and prognostic biomarkers and as tools for therapeutic targets [105].
Circulating miRNA profiles of MPM patients have been studied to identify markers for early detection, differential diagnosis, and prognosis. Several studies have reported increased expression of miR-197-3p, miR-1281, miR-548-3p, miR-20a, miR-625-3p, and miR-34b/c along with downregulation of miR-126, which may be attributed to its tumor suppressor activity [17, 90, 106].
miR-20a and miR-126I have been studied as screening markers [25, 33, 91, 104]: they could be useful in distinguishing healthy individuals who are more or less exposed to asbestos.
Some researchers have demonstrated different expression levels of miR-132 in circulating samples of mesothelioma patients and control subjects exposed to asbestos. The discrimination sensitivity was 86% and the specificity was 61%. When miR-132 was combined with the previously described miR-126, the sensitivity was 77% and the specificity 86% [41].
The results of the studies are somewhat heterogeneous, and this may be primarily attributed to the use of different control groups, small sample sizes, and lack of standardization of circulating microRNA detection methods [24, 43, 45, 73].
CIRCULATING TUMOR DNA (ctDNA) AND EPIGENETIC BIOMARKERS
Recent breakthroughs in medical research have led to new understanding through the study of ctDNA and epigenomic biomarkers.
Circulating free DNA (cfDNA), which comes from healthy and cancerous tissues undergoing apoptosis or necrosis, has shown significant potential in the field of oncology.
On the contrary, circulating tumor DNA (ctDNA), which comes exclusively from cancer cells, carries somatic mutations and accounts for only a small portion of cfDNA [108].
This finding offers a new perspective for previously untreated MPM patients.
In 2018, Hylebos and colleagues conducted a comprehensive analysis of 10 MPM patients using whole-exome sequencers to identify cancer-specific mutations in both germline and tumor DNA [109]. They were able to detect these mutations in serum samples from five treatment-naïve patients, achieving a 60% detection rate. It is interesting to note that no tumor-specific alterations were observed in the cfDNA of patients undergoing chemotherapy. Although ctDNA has the potential to be used as a biomarker of treatment response, further validation and cost-effective technologies will be needed before it can be widely used in routine clinical practice.
Furthermore, epigenetic changes occurring during tumor development have emerged as promising biomarkers detectable in various body fluids.
In asbestos-induced carcinogenesis, the generation of reactive oxygen species (ROS) leads to methylation of gene promoters, orchestrated by polymerase 1 (PARP1) and DNA (cytosine-5) methyltransferase 1 (DNMT1) [110]. Nocchi et al. introduced an innovative approach by combining two epigenetically regulated markers, miR-126 and TM, with SMRP [111]. Indeed, it has been reported that epigenetic mechanisms can silence TM gene expression in MPM tissue, and hypermethylation of the promoter region of miR-126 contributes to its downregulation. Despite a sensitivity rate of 60%, the authors reported that circulating methylated TM DNA effectively differentiated MPM patients from controls with a specificity of 82%. This result complements the performance of miR-126 and SMRP as independent biomarkers for MPM detection [112].
More recently, Guarrera et al. used a genome-wide methylation array to identify distinct methylation patterns at selected CpGs in DNA extracted from white blood cells in a cohort of 163 patients with MPM and 137 controls [113]. This finding promises to shed further light on the epigenetic landscape of MPM and its potential implications for diagnosis and treatment.
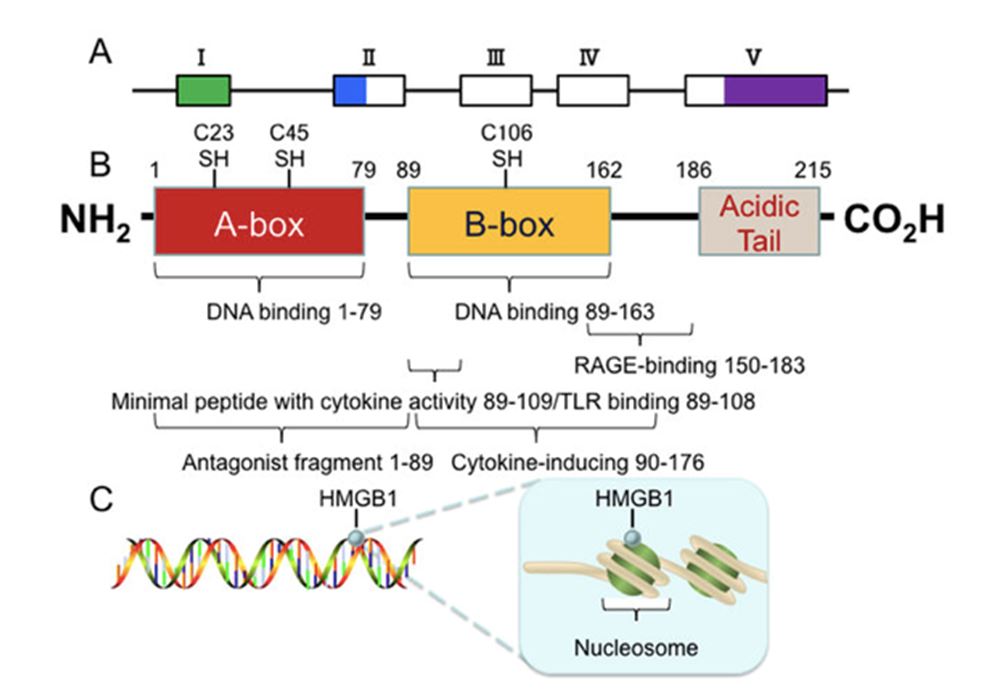
HIGH-MOBILITY GROUP BOX 1 (HMGB1)
HMGB1 is a damage-associated molecular protein that is released in the extracellular space during necrosis [37, 73].
It has been deemed a promising biomarker, with a cytoplasmic hyperacetylated isoform that can be released into the extracellular space, with better results than the nonacetylated form present in the nucleus [25,36]. However, there are conflicting studies in this regard [114]. There are only a few studies showing high marker accuracy in terms of distinguishing between patients with asbestosis and healthy patients with and without asbestos exposure [115]. However, no differences were found when patients with MPM were compared with those with asbestosis, and no studies comparing with other malignancies are available [25, 92].
HMGB1 might also play a role as a potential target for MPM [89].
CONCLUSIONS
As mentioned in this literature review, several scientific efforts were made to test and screen biomarkers derived from the pleural fluid or serum of asbestos-exposed individuals and MPM patients.
Certainly, additional studies will be required to confirm the data reported in this review.
However, the overview that this research offers to the world today is extremely interesting and shows the potential role of biomarkers in the diagnosis and prognosis of MPM, including as potential innovative therapeutic targets.
Reference list
1. Asciak, R.; George, V.; Rahman, N.M. Update on biology and management of mesothelioma. Eur. Respir. Rev. 2021, 30, 200226. [CrossRef] [PubMed]
2. Sauter, J.L.; Dacic, S.; Galateau-Salle, F.; Attanoos, R.L.; Butnor, K.J.; Churg, A.; Husain, A.N.; Kadota, K.; Khoor, A.; Nicholson, A.G.; et al. The 2021 WHO Classification of Tumors of the Pleura: Advances Since the 2015 Classification. J. Thorac. Oncol. 2022, 17, 608–622. [CrossRef] [PubMed]
3. Mlika, M.; Lamzirbi, O.; Limam, M.; Mejri, N.; Saad, S.B.; Chaouch, N.; Miled, K.B.; Marghli, A.; Mezni, F. Clinical and pathological profile of the pleural malignant mesothelioma: A retrospective study about 30 cases. Rev. Pneumol. Clin. 2018, 74, 427–435. [CrossRef] [PubMed]
4. Congedo, M.T.; West, E.C.; Evangelista, J.; Mattingly, A.A.; Calabrese, G.; Sassorossi, C.; Nocera, A.; Chiappetta, M.; Flamini, S.; Abenavoli, L.; et al. The genetic susceptibility in the development of malignant pleural mesothelioma: Somatic and germline variants, clinicopathological features and implication in practical medical/surgical care: A narrative review. J. Thorac. Dis 2023, in press.
5. Moolgavkar, S.H.; Chang, E.T.; Mezei, G.; Mowat, F.S. Epidemiology of Mesothelioma. In Asbestos and Mesothelioma; Testa, J.R., Ed.; Springer International Publishing: Philadelphia, PA, USA, 2017; pp. 43–72.
6. Price, B. Projection of future numbers of mesothelioma cases in the US and the increasing prevalence of background cases: An update based on SEER data for 1975 through 2018. Crit. Rev. Toxicol. 2022, 52, 317–324. [CrossRef] [PubMed]
7. Wen, W.; Xu, D.; Piao, Y.; Li, X. Prognostic value of maximum standard uptake value, metabolic tumour volume, and total lesion glycolysis of 18F-FDG PET/CT in patients with malignant pleural mesothelioma: A systematic review and meta-analysis. Cancer Cell Int. 2022, 22, 60. [CrossRef]
8. Zhang, Y.; Li, R.; Gu, Y.; LiZhu, Y.; Liu, X.; Zhang, S. Clinical, Laboratory, Histological, Radiological, and Metabolic Features and Prognosis of Malignant Pleural Mesothelioma. Medicina 2022, 58, 1874. [CrossRef]
9. Royal College of Physicians (RCP). National Mesothelioma Audit Report 2020 (Audit Period 2016–18); RCP: London, UK, 2020. Available online: www.rcplondon.ac.uk/projects/outputs/national-mesothelioma-audit-report-2020-audit-period-2016-18 (accessed on 2 May 2023).
10. Popat, S.; Baas, P.; Faivre-Finn, C.; Girard, N.; Nicholson, A.G.; Nowak, A.K.; Opitz, I.; Scherpereel, A.; Reck, M.; ESMO Guidelines Committee. Malignant pleural mesothelioma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2022, 33, 129–142. [CrossRef]
11. Shah, R.; Klotz, L.V.; Glade, J. Current Management and Future Perspective in Pleural Mesothelioma. Cancers 2022, 14, 1044. [CrossRef] [PubMed]
12. Vogelzang, N.J.; Rusthoven, J.J.; Symanowski, J.; Denham, C.; Kaukel, E.; Ruffie, P.; Gatzemeier, U.; Boyer, M.; Emri, S.; Manegold, C.; et al. Phase III study of pemetrexed in combination with cisplatin versus cisplatin alone in patients with malignant pleural mesothelioma. J. Clin. Oncol. 2003, 21, 2636–2644. [CrossRef]
13. Zalcman, G.; Mazieres, J.; Margery, J.; Greillier, L.; Audigier-Valette, C.; Moro-Sibilot, D.; Molinier, O.; Corre, R.; Monnet, I.; Gounant, V.; et al. Bevacizumab for newly diagnosed pleural mesothelioma in the Mesothelioma Avastin Cisplatin Pemetrexed Study (MAPS): A randomised, controlled, open-label, phase 3 trial. Lancet 2016, 387, 1405–1414. [CrossRef] [PubMed]
14. Nakajima, E.C.; Vellanki, P.J.; Larkins, E.; Chatterjee, S.; Mishra-Kalyani, P.S.; Bi, Y.; Qosa, H.; Liu, J.; Zhao, H.; Biable, M.; et al. FDA Approval Summary: Nivolumab in Combination with Ipilimumab for the Treatment of Unresectable Malignant Pleural Mesothelioma. Clin. Cancer Res. 2022, 28, 446–451. [CrossRef] [PubMed]
15. Baas, P.; Scherpereel, A.; Nowak, A.K.; Fujimoto, N.; Peters, S.; Tsao, A.S.; Mansfield, A.S.; Popat, S.; Jahan, T.; Antonia, S.; et al. First-line nivolumab plus 15 ipilimumab in unresectable malignant pleural mesothelioma (CheckMate 743): A multicentre, 16 randomised, open-label, phase 3 trial. Lancet 2021, 397, 375–386.17. [CrossRef] [PubMed]
16. Woolhouse, I.; Bishop, L.; Darlison, L.; De Fonseka, D.; Edey, A.; Edwards, J.; Faivre-Finn, C.; Fennell, D.A.; Holmes, S.; Kerr,
K.M.; et al. British Thoracic Society Guideline for the investigation and management of malignant pleural mesothelioma. Thorax 2018, 73 (Suppl. S1), i1–i30. [CrossRef]
17. Scherpereel, A.; Opitz, I.; Berghmans, T.; Psallidas, I.; Glatzer, M.; Rigau, D.; Astoul, P.; B.lükbas, S.; Boyd, J.; Coolen, J.; et al. ERS/ESTS/EACTS/ESTRO guidelines for the management of malignant pleural mesothelioma. Eur. Respir. J. 2020, 55, 1900953. [CrossRef]
18. Pesch, B.; Brüning, T.; Johnen, G.; Casjens, S.; Bonberg, N.; Taeger, D.; Müller, A.;Weber, D.G.; Behrens, T. Biomarker research with prospective study designs for the early detection of cancer. Biochim. Biophys. Acta. 2014, 1844, 874–883. [CrossRef] [PubMed]
19. Liu, X.T.; Dong, X.L.; Zhang, Y.; Fang, P.; Shi, H.Y.; Ming, Z.J. Diagnostic value and safety of medical thoracoscopy for pleural effusion of different causes. World J. Clin. Cases. 2022, 10, 3088–3100. [CrossRef]
20. Dixon, G.; de Fonseka, D.; Maskell, N. Pleural controversies: Image guided biopsy vs. thoracoscopy for undiagnosed pleural effusions? J. Thorac. Dis. 2015, 7, 1041–1051. [PubMed] 21. Zhang, Y.; Li, N.; Li, R.; Gu, Y.; Liu, X.; Zhang, S. Predicting survival for patients with mesothelioma: Development of the PLACE prognostic model. BMC Cancer 2023, 23, 698. [CrossRef] [PubMed]
22. Yeap, B.Y.; De Rienzo, A.; Gill, R.R.; Oster, M.E.; Dao, M.N.; Dao, N.T.; Levy, R.D.; Vermilya, K.; Gustafson, C.E.; Ovsak, G.; et al. Mesothelioma Risk Score: A New Prognostic Pretreatment, Clinical-Molecular Algorithm for Malignant Pleural Mesothelioma. J. Thorac. Oncol. 2021, 16, 1925–1935. [CrossRef]
23. Ahmad, A.; Imran, M.; Ahsan, H. Biomarkers as Biomedical Bioindicators: Approaches and Techniques for the Detection, Analysis, and Validation of Novel Biomarkers of Diseases. Pharmaceutics 2023, 15, 1630. [CrossRef]
24. Zhang, M.; Yan, L.; Lippi, G.; Hu, Z.D. Pleural biomarkers in diagnostics of malignant pleural effusion: A narrative review. Transl. Lung Cancer Res. 2021, 10, 1557–1570. [CrossRef] [PubMed]
25. Schillebeeckx, E.; van Meerbeeck, J.P.; Lamote, K. Clinical utility of diagnostic biomarkers in malignant pleural mesothelioma: A systematic review and meta-analysis. Eur. Respir. Rev. 2021, 30, 210057. [CrossRef] [PubMed]
26. Gao, R.;Wang, F.;Wang, Z.;Wu, Y.; Xu, L.; Qin, Y.; Shi, H.; Tong, Z. Diagnostic value of soluble mesothelin-related peptides in pleural effusion for malignant pleural mesothelioma: An updated meta-analysis. Medicine 2019, 98, e14979. [CrossRef] [PubMed] 27. Cui, A.; Jin, X.G.; Zhai, K.; Tong, Z.H.; Shi, H.Z. Diagnostic values of soluble mesothelin-related peptides for malignant pleural mesothelioma: Updated meta-analysis. BMJ Open 2014, 4, e004145. [CrossRef] 28. Pass, H.I.; Levin, S.M.; Harbut, M.R.; Melamed, J.; Chiriboga, L.; Donington, J.; Huflejt, M.; Carbone, M.; Chia, D.; Goodglick, L.; et al. Fibulin-3 as a blood and effusion biomarker for pleural mesothelioma. N. Engl. J. Med. 2012, 367, 1417–1427; Erratum in N. Engl. J. Med. 2012, 367, 1768. [CrossRef]
29. Fujimoto, N.; Gemba, K.; Asano, M.; Fuchimoto, Y.; Wada, S.; Ono, K.; Ozaki, S.; Kishimoto, T. Hyaluronic acid in the pleural fluid of patients with malignant pleural mesothelioma. Respir. Investig. 2013, 51, 92–97. [CrossRef]
30. Birnie, K.A.; Prêle, C.M.; Musk, A.W.B.; de Klerk, N.; Lee, Y.C.G.; Fitzgerald, D.; Allcock, R.J.N.; Thompson, P.J.; Creaney, J.; Badrian, B.; et al. MicroRNA signatures in malignant pleural mesothelioma effusions. Dis. Markers 2019, 2019, 8628612. [CrossRef]
31. Filiberti, R.; Parodi, S.; Libener, R.; Ivaldi, G.P.; Canessa, P.A.; Ugolini, D.; Bobbio, B.; Marroni, P. Diagnostic value of mesothelin in pleural fluids: Comparison with CYFRA 21-1 and CEA. Med. Oncol. 2013, 30, 543. [CrossRef]
32. Husain, A.N.; Colby, T.V.; Ordóñez, N.G.; Allen, T.C.; Attanoos, R.L.; Beasley, M.B.; Butnor, K.J.; Chirieac, L.R.; Churg, A.M.; Dacic, S.; et al. Guidelines for Pathologic Diagnosis of Malignant Mesothelioma 2017 Update of the Consensus Statement from the International Mesothelioma Interest Group. Arch. Pathol. Lab. Med. 2018, 142, 89–108. [CrossRef]
33. Casjens, S.; Weber, D.G.; Johnen, G.; Raiko, I.; Taeger, D.; Meinig, C.; Moebus, S.; Jöckel, K.-H.; Brüning, T.; Pesch, B. Assessment of potential predictors of calretinin and mesothelin to improve the diagnostic performance to detect malignant mesothelioma: Results from a population-based cohort study. BMJ Open 2017, 7, e017104. [CrossRef]
34. Creaney, J.; Sneddon, S.; Dick, I.M.; Dare, H.; Boudville, N.; Musk, A.W.; Skates, S.J.; Robinson, B.W. Comparison of the diagnostic accuracy of the MSLN gene products, mesothelin and megakaryocyte potentiating factor, as biomarkers for mesothelioma in pleural effusions and serum. Dis. Markers 2013, 35, 119–127. [CrossRef]
35. Yu, Y.; Ryan, B.M.; Thomas, A.; Morrow, B.; Zhang, J.; Kang, Z.; Zingone, A.; Onda, M.; Hassan, R.; Pastan, I.; et al. Elevated
Serum Megakaryocyte Potentiating Factor as a Predictor of Poor Survival in Patients with Mesothelioma and Primary Lung Cancer. J. Appl. Lab. Med. 2018, 3, 166–177. [CrossRef]
36. Napolitano, A.; Antoine, D.J.; Pellegrini, L.; Baumann, F.; Pagano, I.; Pastorino, S.; Goparaju, C.M.; Prokrym, K.; Canino, C.; Pass, H.I.; et al. HMGB1 and Its Hyperacetylated Isoform are Sensitive and Specific Serum Biomarkers to Detect Asbestos Exposure
and to Identify Mesothelioma Patients. Clin. Cancer Res. 2016, 22, 3087–3096. [CrossRef] [PubMed]
37. Creaney, J.; Yeoman, D.; Demelker, Y.; Segal, A.; Musk, A.; Skates, S.J.; Robinson, B.W. Comparison of osteopontin, megakaryocyte potentiating factor, and mesothelin proteins as markers in the serum of patients with malignant mesothelioma. J. Thorac. Oncol. 2008, 3, 851–857. [CrossRef]
38. Arnold, D.T.; De Fonseka, D.; Hamilton, F.W.; Rahman, N.M.; Maskell, N.A. Prognostication and monitoring of mesothelioma using biomarkers: A systematic review. Br. J. Cancer 2017, 116, 731–741. [CrossRef]
39. Pass, H.I.; Alimi, M.; Carbone, M.; Yang, H.; Goparaju, C.M. Mesothelioma Biomarkers: A Review Highlighting Contributions from the Early Detection Research Network. Cancer Epidemiol. Biomark. Prev. 2020, 29, 2524–2540. [CrossRef]
40. Hollevoet, K.; Nackaerts, K.; Gosselin, R.; DeWever,W.; Bosquée, L.; De Vuyst, P.; Germonpré, P.; Kellen, E.; Legrand, C.; Kishi, Y.; et al. Soluble mesothelin, megakaryocyte potentiating factor, and osteopontin as markers of patient response and outcome in mesothelioma. J. Thorac. Oncol. 2011, 6, 1930–1937. [CrossRef]
41. Weber, D.G.; Gawrych, K.; Casjens, S.; Brik, A.; Lehnert, M.; Taeger, D.; Pesch, B.; Kollmeier, J.; Bauer, T.T.; Johnen, G.; et al.
Circulating miR-132-3p as a Candidate Diagnostic Biomarker for Malignant Mesothelioma. Dis. Markers. 2017, 2017, 9280170.
[CrossRef] [PubMed]
42. Munson, P.B.; Hall, E.M.; Farina, N.H.; Pass, H.I.; Shukla, A. Exosomal miR-16-5p as a target for malignant mesothelioma. Sci.
Rep. 2019, 9, 11688. [CrossRef]
43. Cavalleri, T.; Angelici, L.; Favero, C.; Dioni, L.; Mensi, C.; Bareggi, C.; Palleschi, A.; Rimessi, A.; Consonni, D.; Bordini, L.; et al. Plasmatic extracellular vesicle microRNAs in malignant pleural mesothelioma and asbestos-exposed subjects suggest a 2-miRNA signature as potential biomarker of disease. PLoS ONE 2017, 12, e0176680. [CrossRef]
44. Tomasetti, M.; Monaco, F.; Strogovets, O.; Volpini, L.; Valentino, M.; Amati, M.; Neuzil, J.; Santarelli, L. ATG5 as biomarker for early detection of malignant mesothelioma. BMC Res. Notes 2023, 16, 61. [CrossRef] [PubMed]
45. Weber, D.G.; Casjens, S.; Brik, A.; Raiko, I.; Lehnert, M.; Taeger, D.; Gleichenhagen, J.; Kollmeier, J.; Bauer, T.T.; Brüning, T.
Circulating long non-coding RNA GAS5 (growth arrest-specific transcript 5) as a complement marker for the detection of malignant mesothelioma using liquid biopsies. Biomark. Res. 2020, 8, 15. [CrossRef]
46. Matboli, M.; Shafei, A.E.; Ali, M.A.; Gaber, A.I.; Galal, A.; Tarek, O.; Marei, M.; Khairy, E.; El-Khazragy, N.; Anber, N.; et al. Clinical significance of serum DRAM1 mRNA, ARSA mRNA, hsa-miR-2053 and lncRNA-RP1-86D1.3 axis expression in malignant pleural mesothelioma. J. Cell Biochem. 2019, 120, 3203–3211. [CrossRef] [PubMed]
47. Ho, M.; Bera, T.K.; Willingham, M.C.; Onda, M.; Hassan, R.; FitzGerald, D.; Pastan, I. Mesothelin expression in human lung cancer. Clin. Cancer Res. 2007, 13, 1571–1575. [CrossRef] [PubMed]
48. Lv, J.; Li, P. Mesothelin as a biomarker for targeted therapy. Biomark. Res. 2019, 7, 18. [CrossRef]
49. Maeda, M.; Hino, O. Blood tests for asbestos-related mesothelioma. Oncology 2006, 71, 26–31. [CrossRef] [PubMed]
50. Chang, K.; Pai, L.H.; Pass, H.; Pogrebniak, H.W.; Tsao, M.S.; Pastan, I.;Willingham, M.C. Monoclonal antibody K1 reacts with epithelial mesothelioma but not with lung adenocarcinoma. Am. J. Surg. Pathol. 1992, 16, 259–268. [CrossRef]
51. Ordóñez, N.G. Value of mesothelin immunostaining in the diagnosis of mesothelioma. Mod. Pathol. 2003, 16, 192–197. [CrossRef] [PubMed]
52. Ordóñez, N.G. What are the current best immunohistochemical markers for the diagnosis of epithelioid mesothelioma? A review and update. Hum. Pathol. 2007, 38, 1–16. [CrossRef]
53. Kaya, H.; Demir, M.; Taylan, M.; Sezgi, C.; Tanrikulu, A.C.; Yilmaz, S.; Bayram, M.; Kaplan, I.; Senyigit, A. Fibulin-3 as a diagnostic biomarker in patients with malignant mesothelioma. Asian Pac. J. Cancer Prev. 2015, 16, 1403–1407. [CrossRef]
54. Livingstone, I.; Uversky, V.N.; Furniss, D.;Wiberg, A. The Pathophysiological Significance of Fibulin-3. Biomolecules 2020, 10, 1294. [CrossRef]
55. Battolla, E.; Canessa, P.A.; Ferro, P.; Franceschini, M.C.; Fontana, V.; Dessanti, P.; Pinelli, V.; Morabito, A.; Fedeli, F.; Pistillo, M.P.; et al. Comparison of the Diagnostic Performance of Fibulin-3 and Mesothelin in Patients with Pleural Effusions from Malignant Mesothelioma. Anticancer Res. 2017, 37, 1387–1391. [PubMed]
56. Ledda, C.; Senia, P.; Rapisarda, V. Biomarkers for Early Diagnosis and Prognosis of Malignant Pleural Mesothelioma: The Quest Goes on. Cancers 2018, 10, 203. [CrossRef]
57. Creaney, J.; Dick, I.M.; Meniawy, T.M.; Leong, S.L.; Leon, J.S.; Demelker, Y.; Segal, A.; Musk, A.W.; Lee, Y.C.G.; Skates, S.J.; et al. Comparison of fibulin-3 and mesothelin as markers in malignant mesothelioma. Thorax 2014, 69, 895–902. [CrossRef]
58. Joy, R.A.; Vikkath, N.; Ariyannur, P.S. Metabolism and mechanisms of action of hyaluronan in human biology. Drug Metab. Pers. Ther. 2018, 33, 15–32. [CrossRef]
59. Asplund, T.; Versnel, M.A.; Laurent, T.C.; Heldin, P. Human mesothelioma cells produce factors that stimulate the production of hyaluronan by mesothelial cells and fibroblasts. Cancer Res. 1993, 53, 388–392.
60. Creaney, J.; Dick, I.M.; Segal, A.; Musk, A.W.; Robinson, B.W. Pleural effusion hyaluronic acid as a prognostic marker in pleural malignant mesothelioma. Lung Cancer 2013, 82, 491–498. [CrossRef] [PubMed]
61. Smolarz, B.; Durczy ´ nski, A.; Romanowicz, H.; Szyłło, K.; Hogendorf, P. miRNAs in Cancer (Review of Literature). Int. J. Mol. Sci. 2022, 23, 2805. [CrossRef] [PubMed]
62. Liu, L.; Xie, W.; Xue, P.; Wei, Z.; Liang, X.; Chen, N. Diagnostic accuracy and prognostic applications of CYFRA 21-1 in head and neck cancer: A systematic review and meta-analysis. PLoS ONE 2019, 14, e0216561. [CrossRef] [PubMed]
63. Hegmans, J.; Veltman, J.D.; Fung, E.T.; Verch, T.; Glover, C.; Zhang, F.; Allard, W.J.; T’Jampens, D.; Hoogsteden, H.C.; Lambrecht, B.N.; et al. Protein profiling of pleural effusions to identify malignant pleural mesothelioma using SELDI-TOF MS. Technol. Cancer Res. Treat. 2009, 8, 323–332. [CrossRef] [PubMed]
64. Fuhrman, C.; Duche, J.C.; Chouaid, C.; Alsamad, I.A.; Atassi, K.; Monnet, I.; Tillement, J.; Housset, B. Use of tumor markers for differential diagnosis of mesothelioma and secondary pleural malignancies. Clin. Biochem. 2000, 33, 405–410. [CrossRef]
65. Wang, X.F.;Wu, Y.H.;Wang, M.S.;Wang, Y.S. CEA, AFP, CA125, CA153 and CA199 in malignant pleural effusions predict the cause. Asian Pac. J. Cancer Prev. 2014, 15, 363–368. [CrossRef] [PubMed]
66. Mundt, F.; Nilsonne, G.; Arslan, S.; Csür.s, K.; Hillerdal, G.; Yildirim, H.; Metintas, M.; Dobra, K.; Hjerpe, A. Hyaluronan and N-ERC/mesothelin as key biomarkers in a specific two-step model to predict pleural malignant mesothelioma. PLoS ONE 2013, 8, e72030. [CrossRef]
67. Palstrøm, N.B.; Overgaard, M.; Licht, P.; Beck, H.C. Identification of Highly Sensitive Pleural Effusion Protein Biomarkers for Malignant Pleural Mesothelioma by Affinity-Based Quantitative Proteomics. Cancers 2023, 15, 641. [CrossRef] [PubMed]
68. Henderson, D.W.; Reid, G.; Kao, S.C.; van Zandwijk, N.; Klebe, S. Challenges and controversies in the diagnosis of mesothelioma: Part 1. Cytology-only diagnosis, biopsies, immunohistochemistry, discrimination between mesothelioma and reactive mesothelial hyperplasia, and biomarkers. J. Clin. Pathol. 2013, 66, 847–853. [CrossRef] [PubMed]
69. Erber, R.;Warth, A.; Muley, T.; Hartmann, A.; Herpel, E.; Agaimy, A. BAP1 Loss is a Useful Adjunct to Distinguish Malignant
Mesothelioma Including the Adenomatoid-like Variant From Benign Adenomatoid Tumors. Appl. Immunohistochem. Mol. Morphol. 2020, 28, 67–73. [CrossRef]
70. Biancosino, C.; van der Linde, L.I.S.; Sauter, G.; Stellmacher, F.; Krüger, M.; Welker, L. Cytological Diagnostic Procedures in
Malignant Mesothelioma. Adv. Exp. Med. Biol. 2022, 1374, 41–49.
71. Straccia, P.; Magnini, D.; Trisolini, R.; Lococo, F.; Chiappetta, M.; Cancellieri, A. The value of cytology in distinguishing malignant mesothelioma: An institutional experience of 210 cases reclassified according to the International System for Reporting Serous Fluid Cytopathology (ISRSFC). Cytopathology 2022, 33, 77–83. [CrossRef]
72. Wang, J.J.; Yan, L. Serum diagnostic markers for malignant pleural mesothelioma: A narrative review. Transl. Cancer Res. 2022, 11, 4434–4440. [CrossRef]
73. Yeo, D.; Castelletti, L.; van Zandwijk, N.; Rasko, J.E.J. Hitting the Bull’s-Eye: Mesothelin’s Role as a Biomarker and Therapeutic Target for Malignant Pleural Mesothelioma. Cancers 2021, 13, 3932. [CrossRef]
74. Hollevoet, K.; Reitsma, J.B.; Creaney, J.; Grigoriu, B.D.; Robinson, B.W.; Scherpereel, A.; Cristaudo, A.; Pass, H.I.; Nackaerts, K.; Portal, J.A.R.; et al. Serum mesothelin for diagnosing malignant pleural mesothelioma: An individual patient data meta-analysis. J. Clin. Oncol. 2012, 30, 1541–1549. [CrossRef]
75. Hollevoet, K.; Nackaerts, K.; Thimpont, J.; Germonpré, P.; Bosquée, L.; De Vuyst, P.; Legrand, C.; Kellen, E.; Kishi, Y.; Delanghe, J.R.; et al. Diagnostic performance of soluble mesothelin and megakaryocyte potentiating factor in mesothelioma. Am. J. Respir. Crit. Care Med. 2010, 181, 620–625. [CrossRef]
76. Tian, L.; Zeng, R.;Wang, X.; Shen, C.; Lai, Y.;Wang, M.; Che, G. Prognostic significance of soluble mesothelin in malignant pleural mesothelioma: A meta-analysis. Oncotarget 2017, 8, 46425–46435. [CrossRef] [PubMed]
77. Goricar, K.; Kovac, V.; Dodic-Fikfak, M.; Dolzan, V.; Franko, A. Evaluation of soluble mesothelin-related peptides and MSLN genetic variability in asbestos-related diseases. Radiol. Oncol. 2020, 54, 86–95. [CrossRef]
78. Fontana, V.; Pistillo, M.P.; Vigani, A.; Canessa, P.A.; Berisso, G.; Giannoni, U.; Ferro, P.; Franceschini, M.C.; Carosio, R.; Tonarelli, M.; et al. Determination of Mesothelin Levels in Pleural Effusion Does Not Help Predict Survival of Patients With Malignant Pleural Mesothelioma. Anticancer Res. 2019, 39, 5219–5223. [CrossRef]
79. Forest, F.; Patoir, A.; Dal Col, P.; Sulaiman, A.; Camy, F.; Laville, D.; Bayle-Bleuez, S.; Fournel, P.; Habougit, C. Nuclear grading, BAP1, mesothelin and PD-L1 expression in malignant pleural mesothelioma: Prognostic implications. Pathology 2018, 50, 635–641. [CrossRef]
80. Hollevoet, K.; Nackaerts, K.; Thas, O.; Thimpont, J.; Germonpré, P.; De Vuyst, P.; Bosquée, L.; Legrand, C.; Kellen, E.; Kishi, Y.; et al. The effect of clinical covariates on the diagnostic and prognostic value of soluble mesothelin and megakaryocyte potentiating factor. Chest 2012, 141, 477–484. [CrossRef]
81. Katz, S.I.; Roshkovan, L.; Berger, I.; Friedberg, J.S.; Alley, E.W.; Simone, C.B., 2nd; Haas, A.R.; Cengel, K.A.; Sterman, D.H.; Albelda, S.M. Serum soluble mesothelin-related protein (SMRP) and fibulin-3 levels correlate with baseline malignant pleural mesothelioma (PM) tumor volumes but are not useful as biomarkers of response in an immunotherapy trial. Lung Cancer 2021, 154, 5–12. [CrossRef]
82. Grigoriu, B.D.; Chahine, B.; Vachani, A.; Gey, T.; Conti, M.; Sterman, D.H.; Marchandise, G.; Porte, H.; Albelda, S.M.; Scherpereel, A. Kinetics of soluble mesothelin in patients with malignant pleural mesothelioma during treatment. Am. J. Respir. Crit. Care Med. 2009, 179, 950–954. [CrossRef] [PubMed]
83. Creaney, J.; Francis, R.J.; Dick, I.M.; Musk, A.W.; Robinson, B.W.S.; Byrne, M.J.; Nowak, A.K. Serum soluble mesothelin concentrations in malignant pleural mesothelioma: Relationship to tumor volume, clinical stage and changes in tumor burden. Clin. Cancer Res. 2011, 17, 1181–1189. [CrossRef] [PubMed]
84. Wheatley-Price, P.; Yang, B.; Patsios, D.; Patel, D.; Ma, C.; Xu, W.; Leighl, N.; Feld, R.; Cho, B.J.; O’Sullivan, B.; et al. Soluble mesothelin-related Peptide and osteopontin as markers of response in malignant mesothelioma. J. Clin. Oncol. 2010, 28, 3316–3322.
[CrossRef] [PubMed]
85. De Fonseka, D.; Arnold, D.T.; Stadon, L.; Morley, A.; Keenan, E.; Darby, M.; Armstrong, L.; Virgo, P.; Maskell, N.A. A prospective study to investigate the role of serial serum mesothelin in monitoring mesothelioma. BMC Cancer 2018, 18, 199. [CrossRef]
86. Grosso, F.; Mannucci, M.; Ugo, F.; Ferro, P.; Cassinari, M.; Vigani, A.; De Angelis, A.M.; Delfanti, S.; Lia, M.; Guaschino, R.;
et al. Pilot Study to Evaluate Serum Soluble Mesothelin-Related Peptide (SMRP) as Marker for Clinical Monitoring of Pleural
Mesothelioma (PM): Correlation with Modified RECIST Score. Diagnostics 2021, 11, 2015. [CrossRef] [PubMed]
87. Burt, B.M.; Lee, H.-S.; De Rosen, V.L.; Hamaji, M.; Groth, S.S.; Wheeler, T.M.; Sugarbaker, D.J. Soluble Mesothelin-Related Peptides to Monitor Recurrence after Resection of Pleural Mesothelioma. Ann. Thorac. Surg. 2017, 104, 1679–1687. [CrossRef] [PubMed]
88. Roshini, A.; Goparaju, C.; Kundu, S.; Nandhu, M.S.; Longo, S.L.; Longo, J.A.; Chou, J.; Middleton, F.A.; Pass, H.I.; Viapiano, M.S. The extracellular matrix protein fibulin-3/EFEMP1 promotes pleural mesothelioma growth by activation of PI3K/Akt signaling. Front. Oncol. 2022, 12, 1014749. [CrossRef] [PubMed]
89. Wang, Y.; Jiang, Z.; Yan, J.; Ying, S. HMGB1 as a Potential Biomarker and Therapeutic Target for Malignant Mesothelioma. Dis. Markers 2019, 2019, 4183157. [CrossRef] [PubMed]
90. Tomasetti, M.; Staffolani, S.; Nocchi, L.; Neuzil, J.; Strafella, E.; Manzella, N.; Mariotti, L.; Bracci, M.; Valentino, M.; Amati, M.; et al. Clinical significance of circulating miR-126 quantification in malignant mesothelioma patients. Clin. Biochem. 2012, 45,
75–581. [CrossRef]
91. Zhang, L.; Zhou, Q.; Han, Y.Q.; Li, P.; Ouyang, P.H.; Wang, M.Y.; Hu, Z.D. Diagnostic accuracy of circulating miR-126 for malignant pleural mesothelioma: A systematic review and meta-analysis. Transl. Cancer Res. 2021, 10, 1856–1862. [CrossRef]
92. Reid, G.; Johnson, T.G.; van Zandwijk, N. Manipulating microRNAs for the Treatment of Malignant Pleural Mesothelioma: Past, Present and Future. Front. Oncol. 2020, 10, 105. [CrossRef]
93. Birnie, K.A.; Prêle, C.M.; Thompson, P.J.; Badrian, B.; Mutsaers, S.E. Targeting microRNA to improve diagnostic and therapeutic approaches for malignant mesothelioma. Oncotarget 2017, 8, 78193–78207. [CrossRef] [PubMed]
94. Creaney, J.; Yeoman, D.; Musk, A.W.; de Klerk, N.; Skates, S.J.; Robinson, B.W. Plasma versus serum levels of osteopontin and mesothelin in patients with malignant mesothelioma—Which is best? Lung Cancer. 2011, 74, 55–60. [CrossRef] [PubMed]
95. Grigoriu, B.D.; Scherpereel, A.; Devos, P.; Chahine, B.; Letourneux, M.; Lebailly, P.; Grégoire, M.; Porte, H.; Copin, M.-C.; Lassalle, P. Utility of osteopontin and serum mesothelin in malignant pleural mesothelioma diagnosis and prognosis assessment. Clin. Cancer Res. 2007, 13, 2928–2935. [CrossRef] [PubMed]
96. Cristaudo, A.; Foddis, R.; Bonotti, A.; Simonini, S.; Vivaldi, A.; Guglielmi, G.; Ambrosino, N.; Canessa, P.A.; Chella, A.; Lucchi, M.; et al. Comparison between plasma and serum osteopontin levels: Usefulness in diagnosis of epithelial malignant pleural mesothelioma. Int. J. Biol. Markers 2010, 25, 164–170. [CrossRef]
97. Hattori, T.; Iwasaki-Hozumi, H.; Bai, G.; Chagan-Yasutan, H.; Shete, A.; Telan, E.F.; Takahashi, A.; Ashino, Y.; Matsuba, T. Both Full-Length and Protease-Cleaved Products of Osteopontin Are Elevated in Infectious Diseases. Biomedicines 2021, 9, 1006. [CrossRef] [PubMed]
98. Kirschner, M.B.; Pulford, E.; Hoda, M.A.; Rozsas, A.; Griggs, K.; Cheng, Y.Y.; Edelman, J.J.B.; Kao, S.C.; Hyland, R.; Dong, Y.; et al. Fibulin-3 levels in malignant pleural mesothelioma are associated with prognosis but not diagnosis. Br. J. Cancer 2015, 113, 963–969. [CrossRef] [PubMed]
99. Tsim, S.; Alexander, L.; Kelly, C.; Shaw, A.; Hinsley, S.; Clark, S.; Evison, M.; Holme, J.; Cameron, E.J.; Sharma, D.; et al. Serum Proteomics and Plasma Fibulin-3 in Differentiation of Mesothelioma from Asbestos-Exposed Controls and Patients with Other Pleural Diseases. J. Thorac. Oncol. 2021, 16, 1705–1717. [CrossRef] [PubMed]
100. Pei, D.; Li, Y.; Liu, X.; Yan, S.; Guo, X.; Xu, X.; Guo, X. Diagnostic and prognostic utilities of humoral fibulin-3 in malignant pleural mesothelioma: Evidence from a meta-analysis. Oncotarget 2017, 8, 13030–13038. [CrossRef]
101. Raiko, I.; Sander, I.; Weber, D.G.; Raulf-Heimsoth, M.; Gillissen, A.; Kollmeier, J.; Scherpereel, A.; Brüning, T.; Johnen, G. Development of an enzyme-linked immunosorbent assay for the detection of human calretinin in plasma and serum of mesothelioma patients. BMC Cancer 2010, 10, 242. [CrossRef]
102. Blum,W.; Pecze, L.; Felley-Bosco, E.; Schwaller, B. Overexpression or absence of calretinin in mouse primary mesothelial cells
inversely affects proliferation and cell migration. Respir. Res. 2015, 16, 153. [CrossRef] [PubMed]
103. Johnen, G.; Gawrych, K.; Raiko, I.; Casjens, S.; Pesch, B.; Weber, D.G.; Taeger, D.; Lehnert, M.; Kollmeier, J.; Bauer, T.; et al. Calretinin as a blood-based biomarker for mesothelioma. BMC Cancer 2017, 17, 386. [CrossRef]
104. Johnen, G.; Burek, K.; Raiko, I.; Wichert, K.; Pesch, B.; Weber, D.G.; Lehnert, M.; Casjens, S.; Hagemeyer, O.; Taeger, D.; et al. Prediagnostic detection of mesothelioma by circulating calretinin and mesothelin—A case-control comparison nested into a prospective cohort of asbestos-exposed workers. Sci. Rep. 2018, 8, 14321. [CrossRef] [PubMed]
105. Han, Y.Q.; Xu, S.C.; Zheng, W.Q.; Hu, Z.D. Diagnostic value of microRNAs for malignant pleural mesothelioma: A mini-review. Thorac. Cancer 2021, 12, 8–12. [CrossRef]
106. Santarelli, L.; Strafella, E.; Staffolani, S.; Amati, M.; Emanuelli, M.; Sartini, D.; Pozzi, V.; Carbonari, D.; Bracci, M.; Pignotti, E.; et al. Association of MiR-126 with soluble mesothelin-related peptides, a marker for malignant mesothelioma. PLoS ONE 2011, 6, e18232. [CrossRef] [PubMed]
107. Singh, A.; Pruett, N.; Pahwa, R.; Mahajan, A.P.; Schrump, D.S.; Hoang, C.D. MicroRNA-206 suppresses mesothelioma progression via the Ras signaling axis. Mol. Ther. Nucleic Acids 2021, 24, 669–681. [CrossRef] [PubMed]
108. Raei, N.; Safaralizadeh, R.; Latifi-Navid, S. Clinical application of circulating tumor DNA in metastatic cancers. Expert Rev. Mol. Diagn. 2023, 1–12. [CrossRef]
109. Hylebos, M.; de Beeck, K.O.; Pauwels, P.; Zwaenepoel, K.; van Meerbeeck, J.P.; Van Camp, G. Tumor-specific genetic variants can be detected in circulating cell-free DNA of malignant pleural mesothelioma patients. Lung Cancer 2018, 124, 19–22. [CrossRef] [PubMed]
110. Christensen, B.C.; Godleski, J.J.; Marsit, C.J.; Houseman, E.A.; Lopez-Fagundo, C.Y.; Longacker, J.L.; Bueno, R.; Sugarbaker, D.J.; Nelson, H.H.; Kelsey, K.T. Asbestos exposure predicts cell cycle control gene promoter methylation in pleural mesothelioma. Carcinogenesis 2008, 29, 1555–1559. [CrossRef] [PubMed]
111. Nocchi, L.; Tomasetti, M.; Amati, M.; Neuzil, J.; Santarelli, L.; Saccucci, F. Thrombomodulin is silenced in malignant mesothelioma by a poly(ADP-ribose) polymerase-1-mediated epigenetic mechanism. J. Biol. Chem. 2011, 286, 19478–19488. [CrossRef]
112. Santarelli, L.; Staffolani, S.; Strafella, E.; Nocchi, L.; Manzella, N.; Grossi, P.; Bracci, M.; Pignotti, E.; Alleva, R.; Borghi, B.; et al. Combined circulating epigenetic markers to improve mesothelin performance in the diagnosis of malignant mesothelioma. Lung Cancer 2015, 90, 457–464. [CrossRef]
113. Guarrera, S.; Viberti, C.; Cugliari, G.; Allione, A.; Casalone, E.; Betti, M.; Ferrante, D.; Aspesi, A.; Casadio, C.; Grosso, F.; et al. Peripheral Blood DNA Methylation as Potential Biomarker of Malignant Pleural Mesothelioma in Asbestos-Exposed Subjects. J. Thorac. Oncol. 2019, 14, 527–539. [CrossRef]
114. Napolitano, A.; Antoine, D.J.; Pellegrini, L.; Baumann, F.; Pagano, I.; Pastorino, S.; Goparaju, C.M.; Prokrym, K.; Canino, C.; Pass, H.I.; et al. Expression of Concern: HMGB1 and Its Hyperacetylated Isoform are Sensitive and Specific Serum Biomarkers to Detect Asbestos Exposure and to Identify Mesothelioma Patients. Clin. Cancer Res. 2020, 26, 1529. [CrossRef]
115. Ying, S.; Jiang, Z.; He, X.; Yu, M.; Chen, R.; Chen, J.; Ru, G.; Chen, Y.; Chen, W.; Zhu, L.; et al. Serum HMGB1 as a Potential Biomarker for Patients with Asbestos-Related Diseases. Dis. Markers 2017, 2017, 5756102. [CrossRef] [PubMed]
116. Cunningham, G.M.; Roman, M.G.; Flores, L.C.; Hubbard, G.B.; Salmon, A.B.; Zhang, Y.; Gelfond, J.; Ikeno, Y. The paradoxical role of thioredoxin on oxidative stress and aging. Arch. Biochem. Biophys. 2015, 576, 32–38. [CrossRef] [PubMed]
117. Demir, M.; Kaya, H.; Taylan, M.; Ekinci, A.; Yılmaz, S.; Teke, F.; Sezgi, C.; Tanrikulu, A.C.; Meteroglu, F.; Senyigit, A. Evaluation of New Biomarkers in the Prediction of Malignant Mesothelioma in Subjects with Environmental Asbestos Exposure. Lung 2016, 194, 409–417. [CrossRef]
118. Hirayama, N.; Tabata, C.; Tabata, R.; Maeda, R.; Yasumitsu, A.; Yamada, S.; Kuribayashi, K.; Fukuoka, K.; Nakano, T. Pleural effusion VEGF levels as a prognostic factor of malignant pleural mesothelioma. Respir. Med. 2011, 105, 137–142. [CrossRef] [PubMed]
119. Yasumitsu, A.; Tabata, C.; Tabata, R.; Hirayama, N.; Murakami, A.; Yamada, S.; Terada, T.; Iida, S.; Tamura, K.; Fukuoka, K.; et al. Clinical significance of serum vascular endothelial growth factor in malignant pleural mesothelioma. J. Thorac. Oncol. 2010, 5, 479–483. [CrossRef] [PubMed]
120. Cristaudo, A.; Bonotti, A.; Guglielmi, G.; Fallahi, P.; Foddis, R. Serum mesothelin and other biomarkers: What have we learned in the last decade? J. Thorac. Dis. 2018, 10 (Suppl. S2), S353–S359. [CrossRef]
121. Rai, A.J.; Flores, R.M. Association of malignant mesothelioma and asbestos related conditions with ovarian cancer: Shared biomarkers and a possible etiological link? Clin. Chem. Lab. Med. 2011, 49, 5–7. [CrossRef]
122. Muley, T.; Dienemann, H.; Herth, F.J.; Thomas, M.; Meister, M.; Schneider, J. Combination of mesothelin and CEA significantly improves the differentiation between malignant pleural mesothelioma, benign asbestos disease, and lung cancer. J. Thorac. Oncol. 2013, 8, 947–951. [CrossRef]
123. Corradi, M.; Goldoni, M.; Alinovi, R.; Tiseo, M.; Ampollini, L.; Bonini, S.; Carbognani, P.; Casalini, A.; Mutti, A. YKL-40 and mesothelin in the blood of patients with malignant mesothelioma, lung cancer and asbestosis. Anticancer Res. 2013, 33, 5517–5524.
124. Bonotti, A.; Foddis, R.; Landi, S.; Melaiu, O.; De Santi, C.; Giusti, L.; Donadio, E.; Ciregia, F.; Mazzoni, M.R.; Lucacchini, A.; et al. A Novel Panel of Serum Biomarkers for PM Diagnosis. Dis. Markers 2017, 2017, 3510984. [CrossRef] [PubMed]
125. Lu, Z.; Zhang, W.; Huang, K.; Zhu, M.; Gu, X.; Wei, D.; Shi, M.; Chen, Y.; Wang, H. Systematic Review, Meta-Analysis and Bioinformatic Analysis of Biomarkers for Prognosis of Malignant Pleural Mesothelioma. Diagnostics 2022, 12, 2210. [CrossRef] [PubMed]
126. Lagniau, S.; Lamote, K.; van Meerbeeck, J.P.; Vermaelen, K.Y. Biomarkers for early diagnosis of malignant mesothelioma: Do we need another moonshot? Oncotarget 2017, 8, 53751–53762. [CrossRef] [PubMed]